How It Works
AdenoQuick 2.0 Kit
Adenovirus vectors with one or multiple transgenes, replication-deficient or oncolytic vectors, WT or modified fiber, E1/E3/E4 deletion, 2 cloning steps in E. coli
- Most versatile system for adenovirus vector construction
- Four shuttle plasmids:
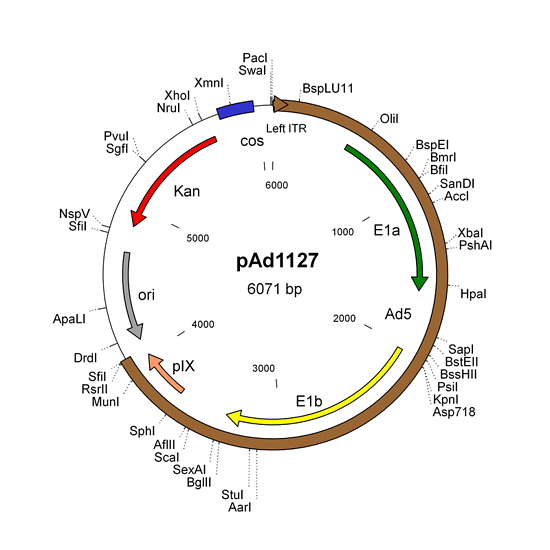
- pAd1127 and derivatives for E1 and pIX genes
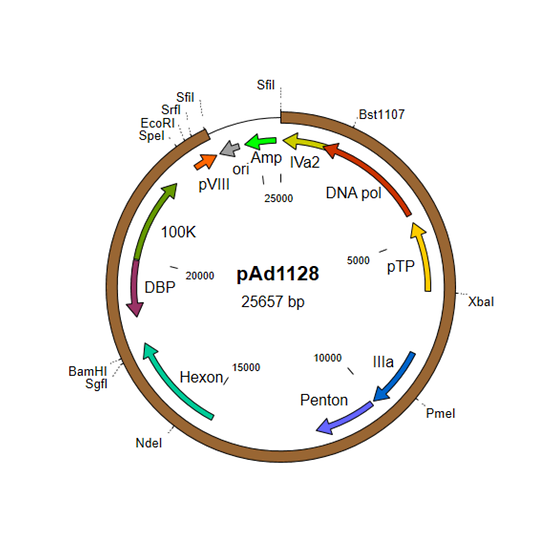
- pAd1128 and derivatives for E2 and late genes
- pAd1129 and derivatives for E3 and fiber genes
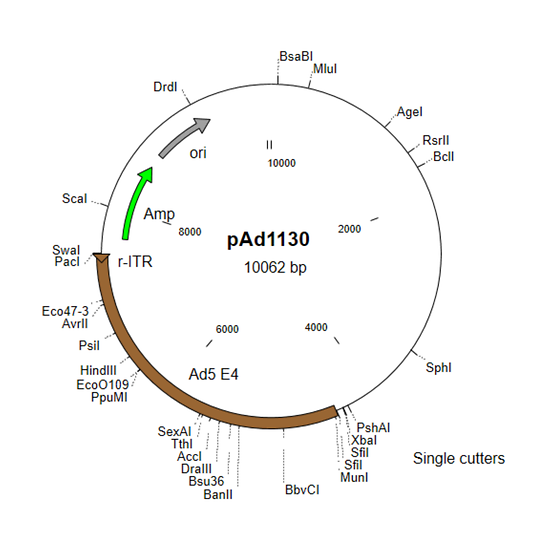
- pAd1130 and derivatives for E4 genes
- Ad5 backbone
- Maximum cargo capacity: 11.2 kb
- Kit contains: 4 shuttle plasmids, reagents for cosmid construction, adenovirus DNA as positive control for virus rescue
Make your kit
Use our Vector Selection Tool to determine which plasmid is most suitable for your application.
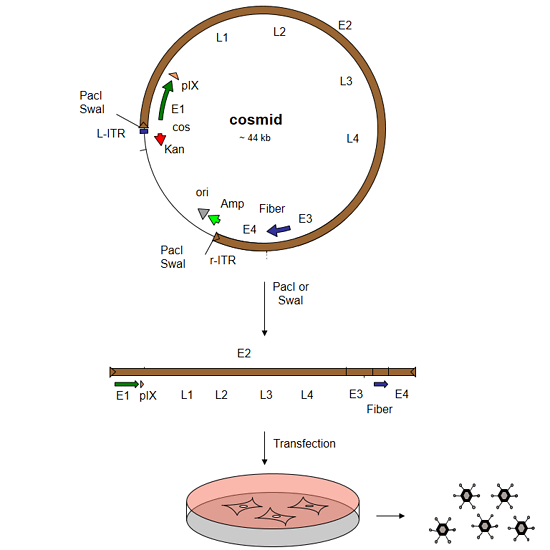
Three steps are needed to rescue a recombinant adenovirus with the AdenoQuick2.0 system:
- Shuttle Plasmid Construction:
Modify one or several of the 4 shuttle plasmids (pAd1127 - pAd1128 - pAd1129 -pAd1130) or their derivatives.- pAd1127 contains the Ad5 left ITR, packaging signal, the E1a and E1b regions, and the pIX coding sequence.
- pAd1128 contains the E2 region (DNA polymerase, terminal protein, DNA-binding protein) and many of the late genes.
- pAd1129 contains the E3 region and fiber gene.
- pAd1130 contains the E4 region and right ITR.
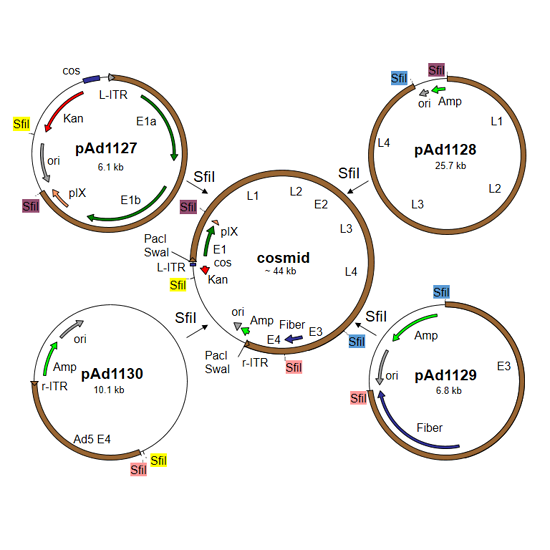
- Cosmid Construction
Construct a cosmid containing the entire genome of your recombinant virus:- Digest the 4 shuttle plasmids with SfiI, and ligate them to each other. The purification of DNA fragments on agarose gel is not necessary.
- Package the ligation products into phage lambda, infect E. coli, and select of ampicilin and kanamycin-resistant clones
- Grow a few colonies, prepare cosmid DNA (= plasmid mini-prep), and confirm their identity by restriction analysis or sequencing
- DNA Transfection
To rescue the virus, you will need to- Linearize the cosmid DNA with PacI or SwaI
- Transfect the linear cosmid DNA into helper cells (e.g. 293 cells)
- Harvest plaques, amplify and confirm virus identity
The following is an up-to-date list of the shuttle plasmids pAd1127, pAd1128, pAd1129, and pAd1130, and their derivatives. Please use our Vector Selection tool to find the plasmid that best suits your needs.
| Description | Cat # | Size |
|---|---|---|
pAd1127Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (WT E1 region) |
QP-04 | 20 µg |
pAd1127-01Shuttle plasmid for the construction of replication-deficient adenovirus vectors using the AdenoQuick2.0 system (E1 deletion) |
QP-05 | 20 µg |
pAd1127-02Shuttle plasmid for the construction of replication-deficient, RCA-free adenovirus vectors using the AdenoQuick2.0 system (E1 deletion) |
QP-06 | 20 µg |
pAd1127-03Shuttle plasmid for the construction of conditionally replicative, oncolytic adenovirus vectors (CrAds) using the AdenoQuick2.0 system |
QP-07 | 20 µg |
pAd1127-04Shuttle plasmid for the construction of conditionally replicative, oncolytic adenovirus vectors (CrAds) using the AdenoQuick2.0 system |
QP-08 | 20 µg |
pAd1127-05Shuttle plasmid for the construction of replication-deficient adenovirus vectors using the AdenoQuick2.0 system (E1 deletion with MCS and SV40 polyA signal) |
QP-23 | 20 µg |
pAd1127-06Shuttle plasmid for the construction of replication-deficient, RCA-free adenovirus vectors using the AdenoQuick2.0 system (E1 deletion, complete packaging signal) |
QP-24 | 20 µg |
pAd1127-07Shuttle plasmid for the construction of replication-deficient adenovirus vectors, with a CMV expression cassette in place of the E1 region, oriented towards the right ITR |
QP-25 | 20 µg |
pAd1127-08Shuttle plasmid for the construction of replication-deficient adenovirus vectors, with a CMV expression cassette in place of the E1 region, oriented towards the left ITR |
QP-26 | 20 µg |
pAd1127-09Shuttle plasmid for the construction of replication-deficient adenovirus vectors, with a RSV expression cassette in place of the E1 region, oriented towards the right ITR |
QP-27 | 20 µg |
pAd1127-10Shuttle plasmid for the construction of replication-deficient adenovirus vectors, with a RSV expression cassette in place of the E1 region, oriented towards the left ITR |
QP-28 | 20 µg |
pAd1127-11Shuttle plasmid for the construction of replication-deficient adenovirus vectors with complete packaging signal and CMV expression cassette in place of the E1 region, oriented towards the right ITR |
QP-29 | 20 µg |
pAd1127-18Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (WT E1 region with 24-bp deletion in CR2) |
QP-37 | 20 µg |
pAd1128Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (WT E2 region and late genes) |
QP-09 | 20 µg |
pAd1129Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (WT E3 region and fiber) |
QP-10 | 20 µg |
pAd1129-01Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (1.9 kb E3 deletion and WT Ad5 fiber) |
QP-11 | 20 µg |
pAd1129-02Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (2.7 kb E3 deletion and WT Ad5 fiber) |
QP-12 | 10 µg |
pAd1129-06Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (2.7 kb E3 deletion and Ad5/35 fiber) |
QP-18 | 20 µg |
pAd1129-10Shuttle plasmid for the construction of adenovirus vectors, with a CMV expression cassette in place of the E3 region, oriented towards the right ITR |
QP-33 | 20 µg |
pAd1129-11Shuttle plasmid for the construction of adenovirus vectors, with a CMV expression cassette in place of the E3 region, oriented towards the left ITR |
QP-34 | 20 µg |
pAd1129-12Shuttle plasmid for the construction of adenovirus vectors, with a RSV expression cassette in place of the E3 region, oriented towards the right ITR |
QP-35 | 20 µg |
pAd1129-13Shuttle plasmid for the construction of adenovirus vectors, with a RSV expression cassette in place of the E3 region, oriented towards the left ITR |
QP-36 | 20 µg |
pAd1129-27Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (2.7 kb E3 deletion and Ad5/3 fiber) |
QP-38 | 20 µg |
pAd1129-34Shuttle plasmid for the construction of adenovirus vectors using the AdenoQuick2.0 system (1.2 kb E3 deletion with Ad5/3 fiber) |
QP-39 | 10 µg |
pAd1130Shuttle vector for the construction of adenovirus vectors using the AdenoQuick2.0 system (WT E4 region) |
QP-13 | 20 µg |
pAd1130-01Shuttle plasmid for constructing adenovirus vectors with the E4 region under the control of a heterologous promoter |
QP-14 | 20 µg |
pAd1130-02Shuttle plasmid for constructing adenovirus vectors with the E4 region under the control of a heterologous promoter with its own TATA box |
QP-15 | 20 µg |
pAd1130-03Shuttle plasmid for constructing adenovirus vectors with a 2.8 kb deletion in the E4 region |
QP-16 | 20 µg |
pAd1130-04Shuttle plasmid for constructing adenovirus vectors with a 1.2 kb deletion in the E4 region |
QP-19 | 20 µg |
Using the AdenoQuick2.0 system to construct your recombinant adenovirus vectors has many advantages:
- The cosmid technology that our system uses is particularly well suited for the cloning of the 36 kb-long adenovirus genome in E coli. Because phage lambda packages DNAs ranging from 39 to 54 kb, the cloning method selects bacterial clones containing full-size adenovirus genomes.
- No homologous recombination event in E. coli is necessary, thus:
- There is no danger for an unpredicted recombination in E. coli that would not be detected by restriction analysis and would cause the DNA to be non-infectious.
- There is no need for the transformation of a recA+ strain (e.g.. BJ5183) for the recombination and the subsequent transfer to a recA endA strain for plasmid preparation. Phage lambda infection can be performed directly into recA endA strains such a DH5a, XL-1 blue or Top10.
- The cosmid construction is very efficient:
- Expression cassettes are linked to a bacterial positive-selection markers (Kanamycin-resistance gene or lambda cos site). Therefore their presence is ensured in the resulting construct. Please note that the Kanamycin-resistance gene is not inserted into the recombinant Ad genome.
- The SfiI restriction sites used in the procedure generate non-symmetrical cohesive ends suitable for directional cloning.
- The cosmid construction via packaging into phage lambda generally produces hundreds of clones, with ~ 100% efficiency. The cosmid construction by electroporation is about 70% efficient.
- Up to 3 µg cosmid DNA per mL bacterial culture can be obtained.
- The method is fast:
- The 4 shuttle plasmids can be modified independently from each other, then assembled together in a single cloning step.
- Plaques usually appear 7 to 10 days after transfection, sometimes as early as 4 days.
- The method is versatile:
- Each of the 4 shuttle plasmids comes in a growing number of versions. The result is several hundred possible configurations for your recombinant virus.
- Two rare-cutting enzymes (PacI and SwaI) are available to linearize the cosmid before transfection into helper cells. Therefore, this method is likely to be useful in a very large number of applications.
- Compared to other techniques that also reconstitute the entire sequence of the recombinant virus in plasmid, AdenoQuick requires less “hands-on” time: given the very high percentage of correct clones obtained using the cosmid system (~100 %), only a limited number of clones needs to be analyzed at small scale (mini preps).
The AdenoQuick2.0 cloning system is ideal for constructing:
- E1-substituted adenovirus vectors, the classical expression adenovirus vector wherein an expression cassette of interest replaces the E1 region.
- E3-substituted adenovirus vectors. This type of vector can be used to avoid the emergence of replication-competent adenoviruses (RCA) during virus amplification in 293 cells. The expression cassette of interest is inserted into the E3 region, while the E1 region is kept empty. If necessary the size of the vector is brought to about 36 kb using stuffer DNA. If the viral vector recombines with the WT E1 region inserted into the 293 cell chromosome, a viral genome will be generated, which will be too long to be packaged, and will not generate viral particles. Please note that there are some restrictions about the nature of the expression cassette that can be inserted into the E3 region.
- E1- and E3-substituted bipartite adenovirus vectors, which contain 2 expression cassettes separated by more than 20 kb DNA: one replacing the E1 region, and the other the E3 region. Examples of applications include:
- Vectors containing your gene of interest in the E1 region, and a reporter gene in the E3 region, or vice-versa
- Vectors containing inducible expression systems
- Vectors expressing a regulator and a regulated gene
- Vectors expressing two genes involved in the same pathway
- E1/E3/E4-deleted adenovirus vectors, with expression cassettes in the E1 or the E3 regions, or both.
- Oncolytic conditionally-replicative adenovirus vectors (CRAd's), wherein a cell-specific promoter drives the expression of E1a, so that the virus replicates only in that cell type. Additional transgenes ("therapeutic", killer, reporter genes) can be inserted in other regions of the virus such as E3.
- Targeted vectors, wherein the Ad5 fiber has been modified so that the virus can infect other cell types
- Vaccine vectors, based on Ad5 or other serotypes such as Ad7, Ad11, Ad26 or Ad35
- Ad5 backbone
- Packaging signal:
- WT (Repeats AI-AVII)
- Partial (Repeats AI-AV)
- E1 region:
- 3.0 kb deletion, corresponding to Ad5 bp 354-3329
- 3.1 kb deletion, corresponding to Ad5 bp 354-3503
- 0.1 kb deletion corresponding to Ad5 bp 354-466
- E3 region:
- WT
- dl309 (0.75 kb deletion corresponding to Ad5 bp 30005-30750 substituted with a 642 bp heterologous DNA)
- 1.9 kb XbaI deletion,corresponding to Ad5 bp 28592-30470
- 2.7 kb BglII deletion, corresponding to Ad5 bp 28133-30818
- Fiber gene:
- WT Ad5
- hybrid Ad5/35
- hybrid Ad5/3
- E4 region:
- WT
- 1.2 kb deletion, corresponding to Ad5 bp 35319-35355 and including ORF1-4
- 2.8 kb deletion, corresponding to Ad5 bp 32830-35649
AdenoQuick2.0 Manual
The AdenoQuick2.0 manual contains all the instructions to construct adenovirus expression vectors using the AdenoQuick2.0 system, including:
- Shuttle plasmid construction
- Cosmid construction
- Virus Rescue
The manual contains also safety guidelines for handling adenovirus vectors, detailed plasmid maps, and troubleshooting guide.